Fresca Melon x Styrofoam Cup #8
RSP 13503
Grower: Team Elite Genetics
General Information
- Sample Name
- S_x_Fresca_#8_20251230
- Accession Date
- December 28, 2025
- Reported Plant Sex
- Female
- Report Type
- Whole-Genome Sequencing
The strain rarity visualization shows how distant the strain is from the other cultivars in the Kannapedia database. The y-axis represents genetic distance, getting farther as you go up. The width of the visualization at any position along the y-axis shows how many strains there are in the database at that genetic distance. So, a common strain will have a more bottom-heavy shape, while uncommon and rare cultivars will have a visualization that is generally shifted towards the top.
Chemical Information
Cannabinoid and terpenoid information provided by the grower.
Cannabinoids
No information provided.
Terpenoids
No information provided.
Genetic Information
- Plant Type
- Type I
File Downloads
The bell curve in the heterozygosity visualization shows the distribution of heterozygosity levels for cannabis cultivars in the Kannapedia database. The green line shows where this particular strain fits within the distribution. Heterozygosity is associated with heterosis (aka hybrid vigor) but also leads to the production of more variable offspring. When plants have two genetically different parents, heterozygosity levels will be higher than if it has been inbred or backcrossed repeatedly.
The ratio of reads mapped to Y-contigs to reads mapped to the whole Cannabis genome (Y-ratios) has been demonstrated to be strongly correlated with plant sex typing. This plot shows the distribution of Y-ratios for all samples in our database which were sequenced with the same method (panel or WGS) as this sample and where this sample falls in the distribution.
This chart represents the Illumina sequence coverage over the Bt/Bd allele. These are the three regions in the cannabis genome that impact THCA, CBDA, CBGA production. Coverage over the Active CBDAS gene is highly correlated with Type II and Type III plants as described by Etienne de Meijer. Coverage over the THCA gene is highly correlated with Type I and Type II plants but is anti-correlated with Type III plants. Type I plants require coverage over the inactive CBDA loci and no coverage over the Active CBDA gene. Lack of coverage over the Active CBDA and Active THCA allele are presumed to be Type IV plants (CBGA dominant). While deletions of entire THCAS and CBDAS genes are the most common Bt:Bd alleles observed, it is possible to have plants with these genes where functional expression of the enzyme is disrupted by deactivating point mutations (Kojoma et al. 2006).
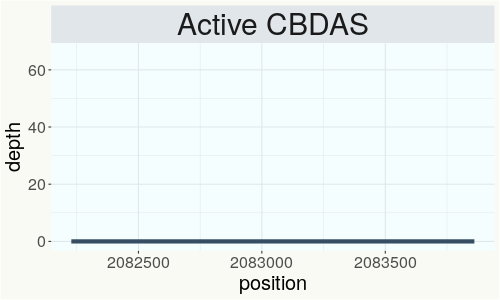
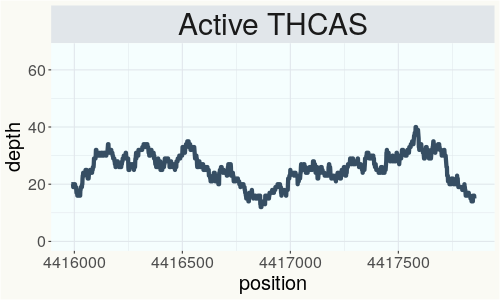
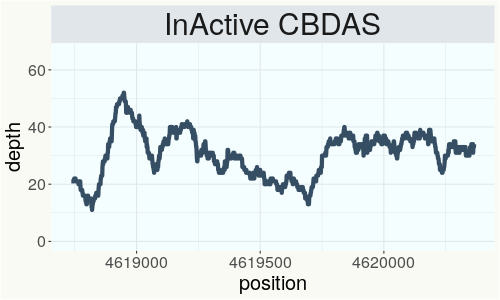
This chart represents the Illumina sequence coverage over the CBCA synthase gene.
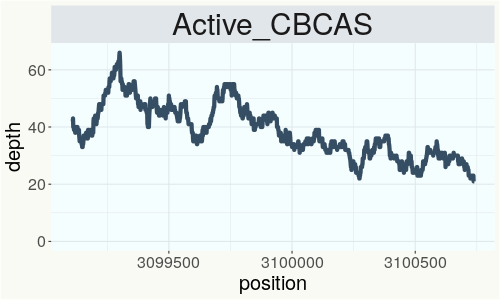
Variants (THCAS, CBDAS, and CBCAS)
Variants (Select Genes of Interest)
| PHL-2 | c.932T>C | p.Leu311Pro | missense variant | moderate | contig2621 | 340210 | T/C | |
| PHL-2 | c.1057A>G | p.Arg353Gly | missense variant | moderate | contig2621 | 340335 | A/G | |
| PHL-2 | c.1096G>A | p.Ala366Thr | missense variant | moderate | contig2621 | 340374 | G/A | |
| PHL-2 | c.2564T>A | p.Phe855Tyr | missense variant | moderate | contig2621 | 342607 | T/A | |
| PHL-2 | c.2578T>A | p.Leu860Ile | missense variant | moderate | contig2621 | 342621 | T/A | |
| PHL-2 | c.2624C>T | p.Ser875Phe | missense variant | moderate | contig2621 | 342667 | C/T | |
| PHL-2 | c.2783G>A | p.Ser928Asn | missense variant | moderate | contig2621 | 342826 | G/A | |
| PHL-2 | c.2830A>C | p.Asn944His | missense variant | moderate | contig2621 | 342873 | A/C |
|
| PHL-2 | c.2933G>T | p.Arg978Leu | missense variant | moderate | contig2621 | 342976 | G/T | |
| PHL-2 | c.2936T>G | p.Val979Gly | missense variant | moderate | contig2621 | 342979 | T/G |
|
| PHL-2 | c.3209A>G | p.Gln1070Arg | missense variant | moderate | contig2621 | 343252 | A/G | |
| PHL-2 | c.3467A>G | p.Gln1156Arg | missense variant | moderate | contig2621 | 343510 | A/G | |
| PKSG-4a | c.617A>G | p.Tyr206Cys | missense variant | moderate | contig700 | 1938028 | A/G |
|
| PKSG-4a |
c.626_628del |
p.Asn209del | disruptive inframe deletion | moderate | contig700 | 1938032 | CAAT/C |
|
| PKSG-4a |
c.1191_1193d |
p.Tyr398del | disruptive inframe deletion | moderate | contig700 | 1938600 | AATT/A |
|
| PKSG-2a | c.1136G>A | p.Arg379His | missense variant | moderate | contig700 | 1944254 | C/T |
|
| PKSG-2a | c.948T>G | p.Asp316Glu | missense variant | moderate | contig700 | 1944442 | A/C |
|
| PKSG-2a | c.945T>G | p.Ser315Arg | missense variant | moderate | contig700 | 1944445 | A/C |
|
| PKSG-2a | c.944G>A | p.Ser315Asn | missense variant | moderate | contig700 | 1944446 | C/T |
|
| PKSG-2a | c.934C>G | p.His312Asp | missense variant | moderate | contig700 | 1944456 | G/C |
|
| PKSG-2a | c.781T>A | p.Leu261Ile | missense variant | moderate | contig700 | 1944609 | A/T | |
| PKSG-2a | c.774G>A | p.Met258Ile | missense variant | moderate | contig700 | 1944616 | C/T |
|
| PKSG-2b | c.67A>T | p.Ile23Phe | missense variant | moderate | contig700 | 1951815 | T/A | |
| PKSG-2b | c.31A>T | p.Thr11Ser | missense variant | moderate | contig700 | 1951851 | T/A | |
| PKSG-4b |
c.535_545del |
p.Ile179fs | frameshift variant | high | contig700 | 2721127 |
CCCCACTCCAAT |
|
| PKSG-4b | c.489delT | p.Phe163fs | frameshift variant | high | contig700 | 2721183 | CA/C | |
| PKSG-4b |
c.353_354ins |
p.Gly119fs | frameshift variant | high | contig700 | 2721319 | T/TGG |
|
| PKSG-4b | c.316+2T>A | splice donor variant & intron variant | high | contig700 | 2723818 | A/T | ||
| PKSG-4b | c.238T>C | p.Phe80Leu | missense variant | moderate | contig700 | 2724197 | A/G |
|
| DXR-2 | c.1319T>C | p.Ile440Thr | missense variant | moderate | contig380 | 285250 | A/G |
|
| DXR-2 | c.431C>G | p.Ala144Gly | missense variant | moderate | contig380 | 287760 | G/C | |
| FAD2-2 | c.196T>C | p.Phe66Leu | missense variant | moderate | contig83 | 1803173 | A/G |
|
| FAD2-2 | c.185C>T | p.Ala62Val | missense variant | moderate | contig83 | 1803184 | G/A |
|
| FAD2-2 | c.172G>T | p.Asp58Tyr | missense variant | moderate | contig83 | 1803197 | C/A |
|
| FAD2-2 | c.161T>A | p.Leu54His | missense variant | moderate | contig83 | 1803208 | A/T |
|
| ELF3 | c.358G>A | p.Gly120Arg | missense variant | moderate | contig97 | 242064 | G/A | |
| ELF3 | c.520A>C | p.Asn174His | missense variant | moderate | contig97 | 242226 | A/C | |
| ELF3 | c.574A>G | p.Asn192Asp | missense variant | moderate | contig97 | 242280 | A/G | |
| ELF3 | c.772A>G | p.Ser258Gly | missense variant | moderate | contig97 | 242478 | A/G |
|
| ELF3 | c.812G>C | p.Gly271Ala | missense variant | moderate | contig97 | 242518 | G/C | |
| ELF3 |
c.1230-2_123 |
splice acceptor variant & intron variant | high | contig97 | 243676 | TAG/T | ||
| ELF3 | c.1466G>A | p.Ser489Asn | missense variant | moderate | contig97 | 244297 | G/A |
|
| ELF3 | c.1630A>G | p.Thr544Ala | missense variant | moderate | contig97 | 244461 | A/G | |
| ELF3 |
c.1803_1805d |
p.His601del | disruptive inframe deletion | moderate | contig97 | 244625 | ACAT/A |
|
| ELF3 | c.1966C>G | p.Pro656Ala | missense variant | moderate | contig97 | 244797 | C/G | |
| ELF3 | c.2141C>G | p.Pro714Arg | missense variant | moderate | contig97 | 244972 | C/G | |
| ELF3 | c.2198G>T | p.Arg733Leu | missense variant | moderate | contig97 | 245029 | G/T | |
| aPT4 | c.97T>C | p.Tyr33His | missense variant | moderate | contig121 | 2828753 | T/C |
|
| aPT4 | c.153A>C | p.Lys51Asn | missense variant | moderate | contig121 | 2828809 | A/C |
|
| aPT1 | c.406A>G | p.Ile136Val | missense variant | moderate | contig121 | 2839605 | A/G | |
| aPT1 | c.629C>T | p.Thr210Ile | missense variant | moderate | contig121 | 2840237 | C/T | |
| HDS-2 |
c.82_93delGT |
p.Val28_Thr3 |
conservative inframe deletion | moderate | contig95 | 1989748 |
CGTAACCGGAAC |
|
| HDS-2 | c.127T>G | p.Ser43Ala | missense variant | moderate | contig95 | 1989794 | T/G |
|
| AAE1-2 | c.331A>G | p.Asn111Asp | missense variant | moderate | contig81 | 209293 | A/G |
|
| AAE1-2 |
c.948_949ins |
p.Asp317fs | frameshift variant | high | contig81 | 209910 | C/CA |
|
| AAE1-2 | c.952delC | p.Gln318fs | frameshift variant | high | contig81 | 209912 | AC/A |
|
| AAE1-2 | c.953A>G | p.Gln318Arg | missense variant | moderate | contig81 | 209915 | A/G |
|
| AAE1-2 | c.955C>T | p.Arg319Cys | missense variant | moderate | contig81 | 209917 | C/T |
|
| AAE1-2 | c.1006A>G | p.Lys336Glu | missense variant | moderate | contig81 | 209968 | A/G |
|
| AAE1-2 | c.1102C>A | p.His368Asn | missense variant | moderate | contig81 | 210064 | C/A |
|
| AAE1-2 | c.1118C>G | p.Thr373Ser | missense variant | moderate | contig81 | 210080 | C/G |
|
| AAE1-2 | c.1415G>A | p.Ser472Asn | missense variant | moderate | contig81 | 210377 | G/A |
|
| AAE1-2 | c.1417A>G | p.Thr473Ala | missense variant | moderate | contig81 | 210379 | A/G |
|
| AAE1-2 | c.1434G>T | p.Glu478Asp | missense variant | moderate | contig81 | 210396 | G/T |
|
| AAE1-2 | c.1541T>C | p.Val514Ala | missense variant | moderate | contig81 | 210503 | T/C |
|
| PHL-1 | c.2623A>G | p.Thr875Ala | missense variant | moderate | contig1439 | 1487174 | T/C |
|
| PHL-1 | c.2551A>G | p.Thr851Ala | missense variant | moderate | contig1439 | 1487246 | T/C | |
| PHL-1 | c.1387A>G | p.Thr463Ala | missense variant | moderate | contig1439 | 1489811 | T/C | |
| Edestin | c.16T>C | p.Ser6Pro | missense variant | moderate | contig850 | 3065274 | A/G |
|
| TFL1 | c.302-1G>A | splice acceptor variant & intron variant | high | contig1636 | 520616 | C/T | ||
| FAD4 | c.121G>T | p.Val41Phe | missense variant | moderate | contig784 | 1690873 | G/T |
|
| HDS-1 | c.1618A>G | p.Ile540Val | missense variant | moderate | contig1891 | 885936 | T/C | |
| HDS-1 | c.1475C>A | p.Pro492Gln | missense variant | moderate | contig1891 | 886162 | G/T |
|
| HDS-1 | c.1393G>A | p.Ala465Thr | missense variant & splice region variant | moderate | contig1891 | 886355 | C/T | |
| HDS-1 | c.1378G>A | p.Val460Ile | missense variant | moderate | contig1891 | 886370 | C/T |
|
| HDS-1 | c.56C>G | p.Ala19Gly | missense variant | moderate | contig1891 | 889336 | G/C | |
| HDS-1 | c.35G>A | p.Cys12Tyr | missense variant | moderate | contig1891 | 889357 | C/T | |
| HDS-1 |
c.-108+1_-10 |
splice donor variant & intron variant | high | contig1891 | 889975 | A/AC |
|
|
| PIE1-2 | c.1872T>A | p.Asp624Glu | missense variant | moderate | contig1460 | 1190252 | A/T | |
| PIE1-2 | c.1289A>G | p.Asp430Gly | missense variant | moderate | contig1460 | 1192109 | T/C |
|
| EMF2 | c.1772A>G | p.Gln591Arg | missense variant | moderate | contig954 | 3059929 | A/G | |
| FT | c.240C>G | p.Asn80Lys | missense variant | moderate | contig1561 | 3124664 | C/G |
|
| PIE1-1 | c.100G>A | p.Glu34Lys | missense variant | moderate | contig1225 | 2277786 | G/A | |
| PIE1-1 | c.773A>G | p.Asn258Ser | missense variant & splice region variant | moderate | contig1225 | 2279897 | A/G | |
| PIE1-1 | c.811T>C | p.Tyr271His | missense variant | moderate | contig1225 | 2279935 | T/C | |
| PIE1-1 | c.815C>T | p.Pro272Leu | missense variant | moderate | contig1225 | 2279939 | C/T | |
| PIE1-1 | c.1222C>G | p.Gln408Glu | missense variant | moderate | contig1225 | 2281482 | C/G | |
| PIE1-1 | c.1394A>G | p.Asp465Gly | missense variant | moderate | contig1225 | 2281654 | A/G | |
| PIE1-1 | c.3614A>G | p.Lys1205Arg | missense variant | moderate | contig1225 | 2285229 | A/G | |
| PIE1-1 | c.3619G>A | p.Val1207Met | missense variant | moderate | contig1225 | 2285234 | G/A |
|
| PIE1-1 | c.6725C>T | p.Ala2242Val | missense variant | moderate | contig1225 | 2289290 | C/T | |
| PIE1-1 | c.6875T>C | p.Leu2292Ser | missense variant | moderate | contig1225 | 2289440 | T/C | |
| GGR | c.704A>T | p.His235Leu | missense variant | moderate | contig2282 | 549696 | A/T |
|
| PKSB-3 | c.1652A>G | p.Glu551Gly | missense variant | moderate | contig93 | 3339759 | A/G |
|
Nearest genetic relatives (All Samples)
- 0.160 RKM-2018-034 (RSP11126)
- 0.161 Rainbow Belts 2.0 (RSP12802)
- 0.168 Fresca Melon x Styrofoam Cup #6 (RSP13398)
- 0.174 Fresca Melon x Pearadise #15 (RSP13399)
- 0.179 Styrofoam Cup X Cookie Dough #2 (RSP13104)
- 0.186 Pearadise x Fuzzy Melon #7 (RSP13103)
- 0.187 SHERBERT (RSP11355)
- 0.189 Pearadise x Cookie Dough #13 (RSP13105)
- 0.190 Casco Kush (RSP11167)
- 0.194 Pure Power Plant (RSP11265)
- 0.199 Domnesia (RSP11184)
- 0.199 Gorilla Cookies (RSP11231)
- 0.203 Deadhead OG (RSP11463)
- 0.203 East Coast Sour Diesel (RSP12922)
- 0.204 NSPM_x_NSPM (RSP11487)
- 0.206 Styrofoam Cup X Sh3rb3t #2 (RSP13102)
- 0.206 MENDO BREATH (RSP11242)
- 0.206 Pearadise x Cookie Dough #11 (RSP13106)
- 0.207 Runtz (RO3) (RSP13306)
- 0.207 HM Buds (RSP13128)
Most genetically distant strains (All Samples)
- 0.482 Big Red (RSP13217)
- 0.471 Cherry Blossom (RSP11328)
- 0.460 Jamaican Lion (RSP12916)
- 0.458 Cherry Blossom (RSP11323)
- 0.453 Tanao Sri (46) (RSP11486)
- 0.446 Cherry Blossom (RSP11309)
- 0.436 JL yellow (RSP11075)
- 0.435 Jamaican Lion (RSP12917)
- 0.431 JL 3rd Gen Mother (RSP11214)
- 0.428 Cherry Blossom (RSP11301)
- 0.426 Jamaican Lion (RSP12915)
- 0.425 Cherry Blossom (RSP11318)
- 0.421 CBD2 V.2 (RSP12669)
- 0.415 AVIDEKEL 2.0 (RSP11174)
- 0.411 Unknown-(Cherry Wine)-(001) (RSP11268)
- 0.410 Unknown-(Cherry Wine)-(003) (RSP11270)
- 0.407 Cherry Blossom (RSP11298)
- 0.407 Cherry Blossom (RSP11325)
- 0.406 Jamaican Lion (RSP12913)
- 0.405 JL 4th Gen 5 (RSP11199)
Blockchain Registration Information
- Transaction ID
-
RSP13503-mgc-ots
-certificate - SHASUM Hash
-
81bb28d9dd19a0be1f9796653bcbb833 a57f307126873899 15790e65960528a7