#11 Janis
RSP 11654
Grower: Kevin McKernan
General Information

- Accession Date
- September 23, 2020
- Reported Plant Sex
- not reported
- Report Type
- CannSNP90
The strain rarity visualization shows how distant the strain is from the other cultivars in the Kannapedia database. The y-axis represents genetic distance, getting farther as you go up. The width of the visualization at any position along the y-axis shows how many strains there are in the database at that genetic distance. So, a common strain will have a more bottom-heavy shape, while uncommon and rare cultivars will have a visualization that is generally shifted towards the top.
Chemical Information
Cannabinoid and terpenoid information provided by the grower.
Cannabinoids
No information provided.
Terpenoids
No information provided.
Genetic Information
- Plant Type
- Type I
File Downloads
This chart represents the Log-R Ratio (LRR) over variants in the region of the THCA synthase gene. A high correlation between the LRR and samples with a known deletion in THCA synthase indicates the THCA region is deleted and a low correlation indicates it is intact.
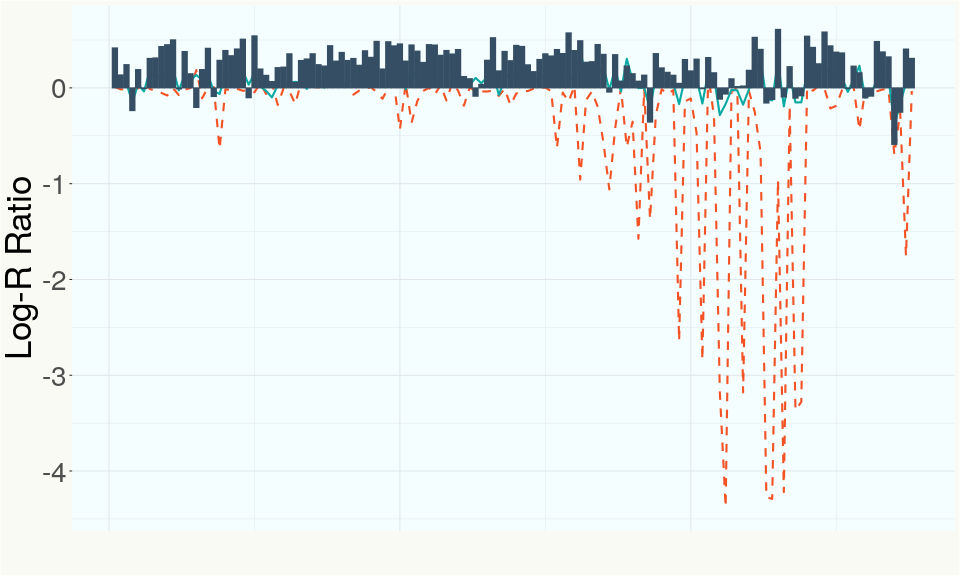
This chart represents the Log-R Ratio (LRR) over variants in the region of the CBDA synthase gene. A high correlation between the LRR and samples with a known deletion in CBDA synthase indicates the CBDA region is deleted and a low correlation indicates it is intact.
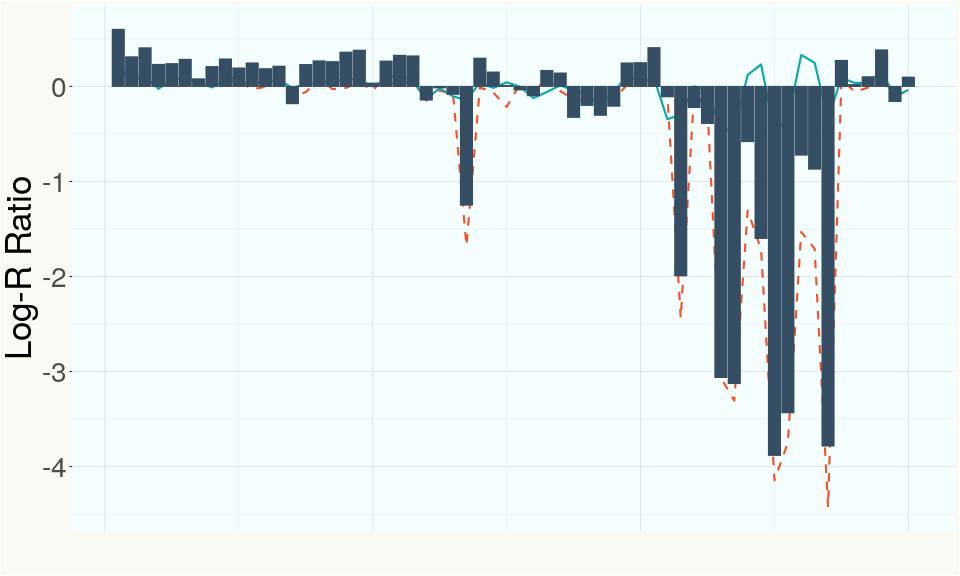
This chart represents the Log-R Ratio (LRR) over variants in the region of the CBCA synthase gene. A high correlation between the LRR and samples with a known deletion in CBCA synthase indicates the CBCA region is deleted and a low correlation indicates it is intact.
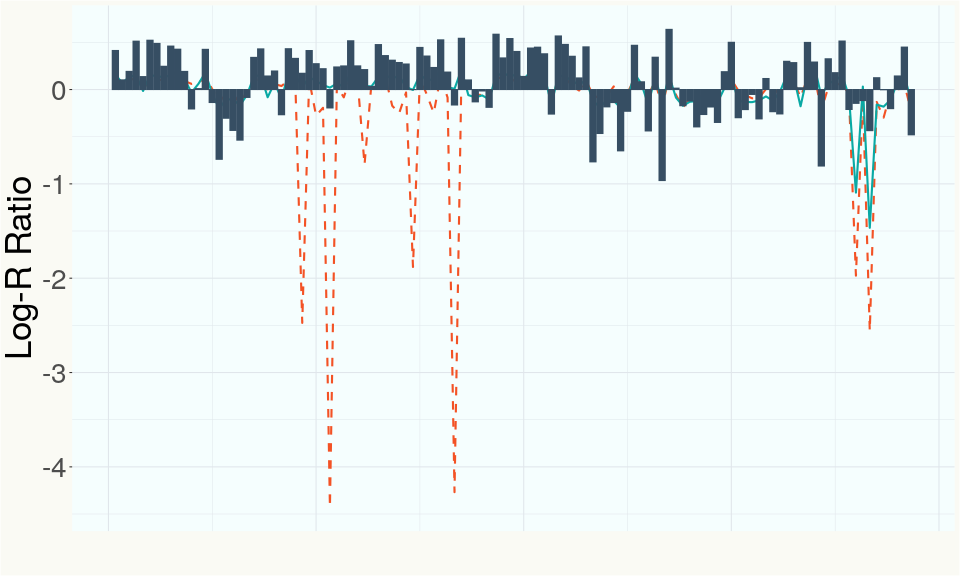
This chart represents the Log-R Ratio (LRR) over variants in the Y-contigs. A high correlation between the LRR and samples which are known Females indicates these Y-contigs are deleted in this sample and a low correlation indicates that the Y-contigs are not deleted and is likely Male.
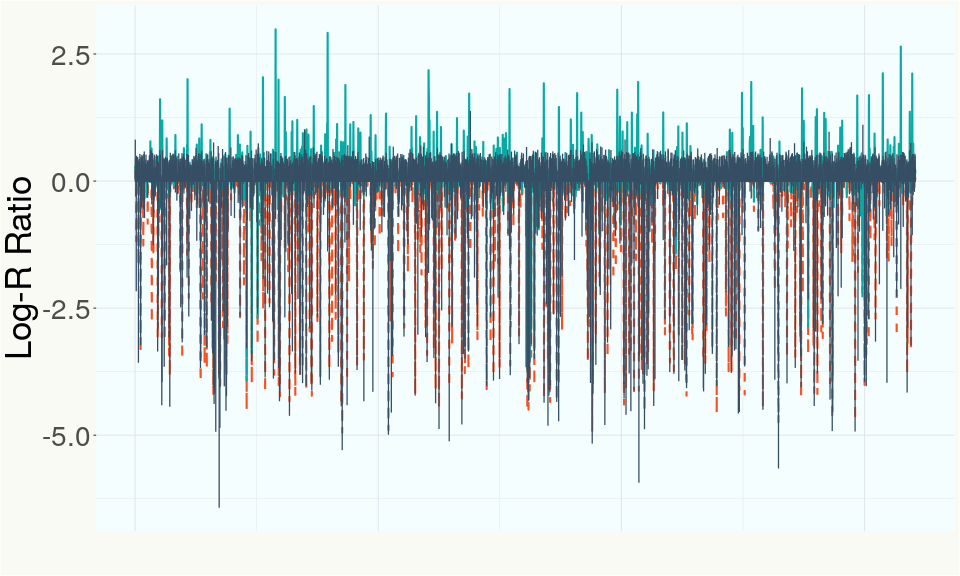
Summary of Deletions
THCAS
- Correlation:
- 0.42
- Call:
- intact
CBDAS
- Correlation:
- 0.98
- Call:
- deleted
CBCAS
- Correlation:
- 0.24
- Call:
- intact
Plant Sex
- Correlation:
- 0.93
- Call:
- female
Variants (THCAS, CBDAS, and CBCAS)
| THCAS | c.749C>A | p.Ala250Asp | missense variant | moderate | contig741 | 4417079 | G/T | |
| THCAS | c.424G>A | p.Val142Ile | missense variant | moderate | contig741 | 4417404 | C/T | |
| THCAS | c.385G>A | p.Val129Ile | missense variant | moderate | contig741 | 4417443 | C/T | |
| THCAS | c.355A>T | p.Met119Leu | missense variant | moderate | contig741 | 4417473 | T/A | |
| THCAS | c.187A>C | p.Ile63Leu | missense variant | moderate | contig741 | 4417641 | T/G |
Variants (Select Genes of Interest)
| PKSA-3b | c.298G>A | p.Ala100Thr | missense variant | moderate | contig705 | 2271640 | C/T | |
| EMF1-2 | c.634G>C | p.Val212Leu | missense variant | moderate | contig885 | 734 | G/C | |
| EMF1-2 | c.710A>C | p.His237Pro | missense variant | moderate | contig885 | 810 | A/C | |
| EMF1-2 | c.1384A>C | p.Lys462Gln | missense variant | moderate | contig885 | 2270 | A/C | |
| PHL-2 | c.44G>A | p.Arg15Lys | missense variant | moderate | contig2621 | 337613 | G/A | |
| PHL-2 | c.932T>C | p.Leu311Pro | missense variant | moderate | contig2621 | 340210 | T/C | |
| PHL-2 | c.1057A>G | p.Arg353Gly | missense variant | moderate | contig2621 | 340335 | A/G | |
| PHL-2 | c.1096G>A | p.Ala366Thr | missense variant | moderate | contig2621 | 340374 | G/A | |
| PHL-2 | c.2564T>A | p.Phe855Tyr | missense variant | moderate | contig2621 | 342607 | T/A | |
| PHL-2 | c.2578T>A | p.Leu860Ile | missense variant | moderate | contig2621 | 342621 | T/A | |
| PHL-2 | c.2624C>T | p.Ser875Phe | missense variant | moderate | contig2621 | 342667 | C/T | |
| PHL-2 | c.2783G>A | p.Ser928Asn | missense variant | moderate | contig2621 | 342826 | G/A | |
| PHL-2 | c.2933G>T | p.Arg978Leu | missense variant | moderate | contig2621 | 342976 | G/T | |
| PHL-2 | c.3209A>G | p.Gln1070Arg | missense variant | moderate | contig2621 | 343252 | A/G | |
| PHL-2 | c.3467A>G | p.Gln1156Arg | missense variant | moderate | contig2621 | 343510 | A/G | |
| PKSG-4a | c.1000T>C | p.Tyr334His | missense variant | moderate | contig700 | 1938411 | T/C | |
| PKSG-2a | c.1117A>G | p.Ile373Val | missense variant | moderate | contig700 | 1944273 | T/C | |
| PKSG-2a | c.995C>T | p.Ser332Phe | missense variant | moderate | contig700 | 1944395 | G/A | |
| PKSG-2a | c.224A>G | p.Lys75Arg | missense variant | moderate | contig700 | 1945166 | T/C | |
| PKSG-2b | c.1117A>G | p.Ile373Val | missense variant | moderate | contig700 | 1950521 | T/C | |
| PKSG-4b | c.523C>T | p.His175Tyr | missense variant | moderate | contig700 | 2721150 | G/A | |
| PKSG-4b | c.489delT | p.Phe163fs | frameshift variant | high | contig700 | 2721183 | CA/C | |
| PKSG-4b | c.316+2T>A | splice donor variant & intron variant | high | contig700 | 2723818 | A/T | ||
| DXR-2 | c.431C>G | p.Ala144Gly | missense variant | moderate | contig380 | 287760 | G/C | |
| OAC-2 | c.220A>G | p.Ile74Val | missense variant | moderate | contig931 | 110019 | T/C | |
| OAC-1 | c.220A>G | p.Ile74Val | missense variant | moderate | contig931 | 118144 | T/C | |
| ELF3 | c.358G>A | p.Gly120Arg | missense variant | moderate | contig97 | 242064 | G/A | |
| ELF3 | c.520A>C | p.Asn174His | missense variant | moderate | contig97 | 242226 | A/C | |
| ELF3 | c.752G>T | p.Gly251Val | missense variant | moderate | contig97 | 242458 | G/T | |
| ELF3 | c.812G>C | p.Gly271Ala | missense variant | moderate | contig97 | 242518 | G/C | |
| ELF3 |
c.1230-2_123 |
splice acceptor variant & intron variant | high | contig97 | 243676 | TAG/T | ||
| ELF3 | c.1630A>G | p.Thr544Ala | missense variant | moderate | contig97 | 244461 | A/G | |
| ELF3 | c.1966C>G | p.Pro656Ala | missense variant | moderate | contig97 | 244797 | C/G | |
| ELF3 | c.2141C>G | p.Pro714Arg | missense variant | moderate | contig97 | 244972 | C/G | |
| ELF3 | c.2198delG | p.Arg733fs | frameshift variant | high | contig97 | 245028 | CG/C | |
| aPT1 | c.406A>G | p.Ile136Val | missense variant | moderate | contig121 | 2839605 | A/G | |
| PHL-1 | c.2551A>G | p.Thr851Ala | missense variant | moderate | contig1439 | 1487246 | T/C | |
| PHL-1 | c.1387A>G | p.Thr463Ala | missense variant | moderate | contig1439 | 1489811 | T/C | |
| TFL1 | c.302-1G>A | splice acceptor variant & intron variant | high | contig1636 | 520616 | C/T | ||
| TFL1 | c.47_48dupAT | p.Val17fs | frameshift variant | high | contig1636 | 521258 | C/CAT | |
| HDS-1 | c.1618A>G | p.Ile540Val | missense variant | moderate | contig1891 | 885936 | T/C | |
| HDS-1 | c.56C>G | p.Ala19Gly | missense variant | moderate | contig1891 | 889336 | G/C | |
| HDS-1 | c.35G>A | p.Cys12Tyr | missense variant | moderate | contig1891 | 889357 | C/T | |
| PIE1-2 | c.6653A>G | p.Asn2218Ser | missense variant | moderate | contig1460 | 1184434 | T/C | |
| PIE1-2 | c.6636T>G | p.Asp2212Glu | missense variant | moderate | contig1460 | 1184451 | A/C | |
| PIE1-2 | c.5932A>G | p.Ile1978Val | missense variant | moderate | contig1460 | 1185552 | T/C | |
| PIE1-2 | c.5132T>C | p.Ile1711Thr | missense variant | moderate | contig1460 | 1186607 | A/G | |
| PIE1-2 |
c.2083_2085d |
p.Val695del | conservative inframe deletion | moderate | contig1460 | 1189954 | GGAC/G | |
| PIE1-2 | c.2072A>G | p.His691Arg | missense variant | moderate | contig1460 | 1189968 | T/C | |
| PIE1-2 | c.1916T>C | p.Val639Ala | missense variant | moderate | contig1460 | 1190124 | A/G | |
| PIE1-2 | c.1872T>A | p.Asp624Glu | missense variant | moderate | contig1460 | 1190252 | A/T | |
| PIE1-2 | c.1630G>C | p.Ala544Pro | missense variant | moderate | contig1460 | 1191600 | C/G | |
| PIE1-2 | c.1156T>G | p.Trp386Gly | missense variant | moderate | contig1460 | 1192242 | A/C | |
| PIE1-2 | c.1117C>G | p.Gln373Glu | missense variant | moderate | contig1460 | 1192281 | G/C | |
| PIE1-2 | c.1093G>A | p.Gly365Ser | missense variant | moderate | contig1460 | 1192305 | C/T | |
| PIE1-2 | c.982G>A | p.Glu328Lys | missense variant | moderate | contig1460 | 1192416 | C/T | |
| PIE1-2 | c.710C>T | p.Pro237Leu | missense variant | moderate | contig1460 | 1193804 | G/A | |
| PIE1-2 | c.706T>C | p.Tyr236His | missense variant | moderate | contig1460 | 1193808 | A/G | |
| PIE1-2 | c.637T>A | p.Ser213Thr | missense variant | moderate | contig1460 | 1194421 | A/T | |
| PIE1-2 | c.349C>T | p.Pro117Ser | missense variant & splice region variant | moderate | contig1460 | 1195017 | G/A | |
| EMF2 | c.1772A>G | p.Gln591Arg | missense variant | moderate | contig954 | 3059929 | A/G | |
| EMF1-1 | c.470C>A | p.Ser157Tyr | missense variant | moderate | contig883 | 269959 | C/A | |
| EMF1-1 | c.605A>T | p.His202Leu | missense variant | moderate | contig883 | 270210 | A/T | |
| FLD | c.2981T>C | p.Met994Thr | missense variant | moderate | contig1450 | 2044012 | A/G | |
| FLD | c.2976A>C | p.Gln992His | missense variant | moderate | contig1450 | 2044017 | T/G | |
| FLD | c.2964C>A | p.Asp988Glu | missense variant | moderate | contig1450 | 2044029 | G/T | |
| FLD | c.2929T>C | p.Phe977Leu | missense variant | moderate | contig1450 | 2044103 | A/G | |
| FLD | c.125G>A | p.Ser42Asn | missense variant | moderate | contig1450 | 2047909 | C/T | |
| PKSA-3a | c.298G>A | p.Ala100Thr | missense variant | moderate | contig1357 | 1144811 | G/A | |
| PIE1-1 | c.742T>A | p.Ser248Thr | missense variant | moderate | contig1225 | 2279320 | T/A | |
| PIE1-1 | c.773A>G | p.Asn258Ser | missense variant & splice region variant | moderate | contig1225 | 2279897 | A/G | |
| PIE1-1 | c.811T>C | p.Tyr271His | missense variant | moderate | contig1225 | 2279935 | T/C | |
| PIE1-1 | c.815C>T | p.Pro272Leu | missense variant | moderate | contig1225 | 2279939 | C/T | |
| PIE1-1 | c.1222C>G | p.Gln408Glu | missense variant | moderate | contig1225 | 2281482 | C/G | |
| PIE1-1 | c.1394A>G | p.Asp465Gly | missense variant | moderate | contig1225 | 2281654 | A/G | |
| PIE1-1 | c.1454T>C | p.Val485Ala | missense variant | moderate | contig1225 | 2281714 | T/C | |
| PIE1-1 |
c.1548_1549i |
p.Gln516_Glu |
conservative inframe insertion | moderate | contig1225 | 2281807 | A/AGAT | |
| PIE1-1 | c.2018T>C | p.Val673Ala | missense variant | moderate | contig1225 | 2283633 | T/C | |
| PIE1-1 | c.2174A>G | p.His725Arg | missense variant | moderate | contig1225 | 2283789 | A/G | |
| PIE1-1 |
c.2185_2187d |
p.Val729del | conservative inframe deletion | moderate | contig1225 | 2283796 | AGTC/A | |
| PIE1-1 | c.3869A>C | p.Asn1290Thr | missense variant | moderate | contig1225 | 2285484 | A/C | |
| PIE1-1 | c.5234T>C | p.Ile1745Thr | missense variant | moderate | contig1225 | 2287152 | T/C | |
| PIE1-1 | c.6041T>C | p.Met2014Thr | missense variant | moderate | contig1225 | 2288214 | T/C | |
| PIE1-1 | c.6738G>T | p.Glu2246Asp | missense variant | moderate | contig1225 | 2289303 | G/T |
Nearest genetic relatives (All Samples)
- 0.001 #12 Donna (RSP11655)
- 0.136 Wedding Cake x MAC (RSP11464)
- 0.138 Thank You Jerry (RSP11459)
- 0.138 #4 Gina (RSP11647)
- 0.140 JL Cross 78 (RSP11579)
- 0.140 #2 Willie (RSP11645)
- 0.140 JL Cross 36 (RSP11537)
- 0.141 UP Wendigo (RSP11261)
- 0.145 Herijuana (RSP11181)
- 0.145 #8 Garcia (RSP11651)
- 0.146 Domnesia (RSP11184)
- 0.146 #5 Football (RSP11648)
- 0.147 JL Cross 56 (RSP11557)
- 0.147 JL Cross 77 (RSP11578)
- 0.148 JL Cross 34 (RSP11535)
- 0.148 JL Cross 84 (RSP11585)
- 0.149 #6 Gloria (RSP11649)
- 0.149 JL Cross 80 (RSP11581)
- 0.151 #7 Gladys (RSP11650)
- 0.151 JL Cross 25 (RSP11526)
Most genetically distant strains (All Samples)
- 0.243 Feral (RSP11205)
- 0.236 CS (RSP11208)
- 0.231 Fedora 17 (RSP11203)
- 0.230 Tiborszallasie (RSP11210)
- 0.227 80E (RSP11212)
- 0.226 Carmagnola/USO 31 (RSP11204)
- 0.226 Feral (RSP11206)
- 0.224 Carmaleonte (RSP11207)
- 0.222 80E (RSP11213)
- 0.218 80E (RSP11211)
- 0.216 Eletta Campana (RSP11209)
- 0.203 JL 4th Gen 6 (RSP11200)
- 0.202 JL 3rd Gen Father (RSP11196)
- 0.196 Swiss Dream (RSP11408)
- 0.196 Unknown-(Cherry Wine)-(004) (RSP11271)
- 0.196 Tahoe OG (RSP11189)
- 0.195 Goomendaze (RSP11462)
- 0.191 JL Cross 33 (RSP11534)
- 0.191 Red Eye OG (RSP11190)
- 0.190 Chematonic (Cannatonic x Chemdawg) (RSP11394)
Blockchain Registration Information
- Transaction ID
-
70e9a7dc70784475
7bc0fed1eb76a306 69c7203ea834fb1c 84c7b1f0ae6a09b7 - Stamping Certificate
- Download PDF (39.4 KB)
- SHASUM Hash
-
4057fa34105e3414e903644ea0397020 59152cb538044899 05e8c6942030fedf